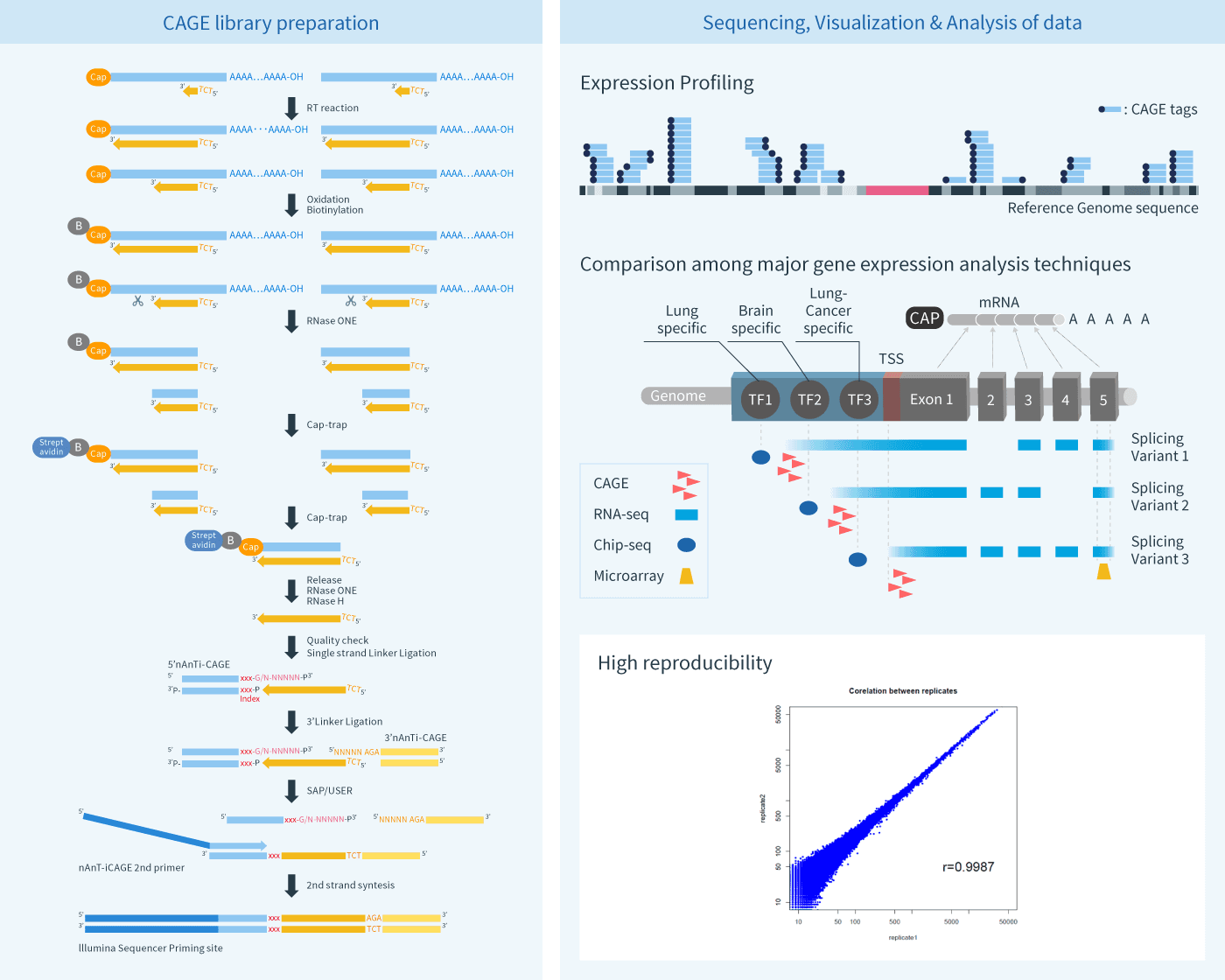
Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage | PNAS

RECLU: a pipeline to discover reproducible transcriptional start sites and their alternative regulation using capped analysis of gene expression (CAGE) | BMC Genomics | Full Text

MAPCap allows high-resolution detection and differential expression analysis of transcription start sites | Nature Communications
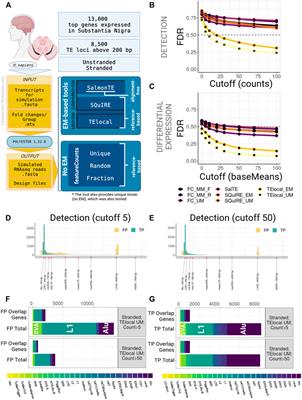
Frontiers | Transcription start site signal profiling improves transposable element RNA expression analysis at locus-level
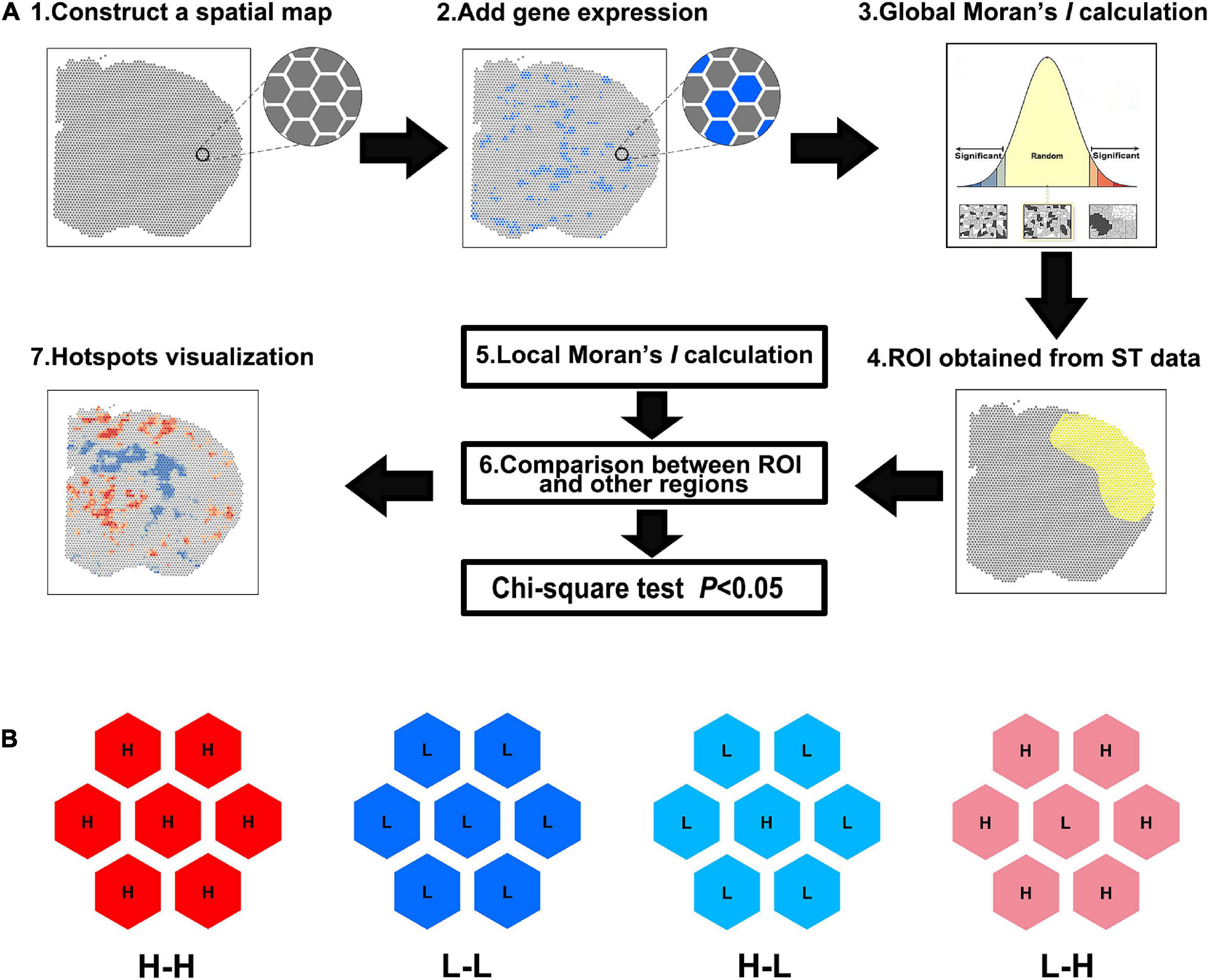
Frontiers | Detection of differentially expressed genes in spatial transcriptomics data by spatial analysis of spatial transcriptomics: A novel method based on spatial statistics

Predictive features of gene expression variation reveal mechanistic link with differential expression | Molecular Systems Biology

CAGE-Seq Reveals that HIV-1 Latent Infection Does Not Trigger Unique Cellular Responses in a Jurkat T Cell Model | Journal of Virology
Dedicated transcriptomics combined with power analysis lead to functional understanding of genes with weak phenotypic changes in knockout lines | PLOS Computational Biology

Development of NET-CAGE a, Schematic of nascent RNA isolation in the... | Download Scientific Diagram
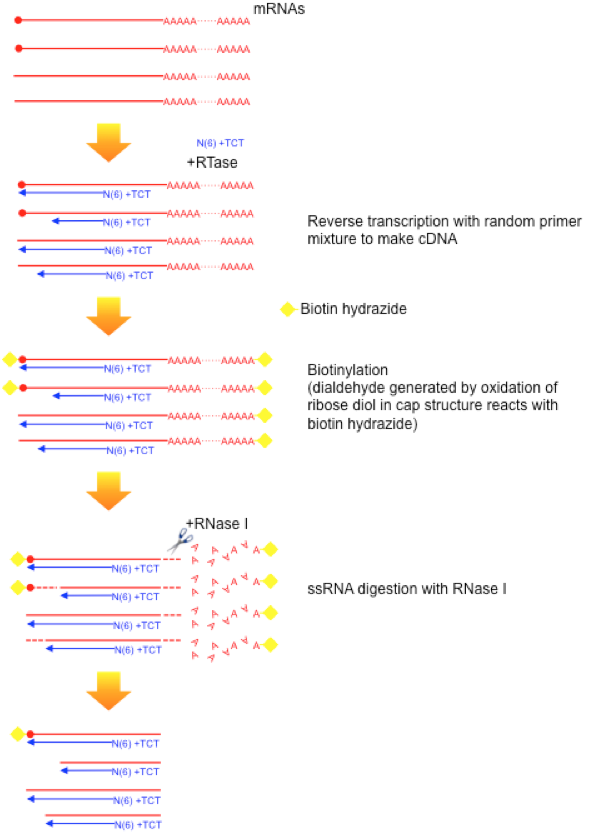
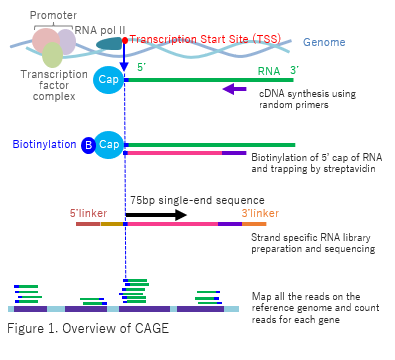
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター
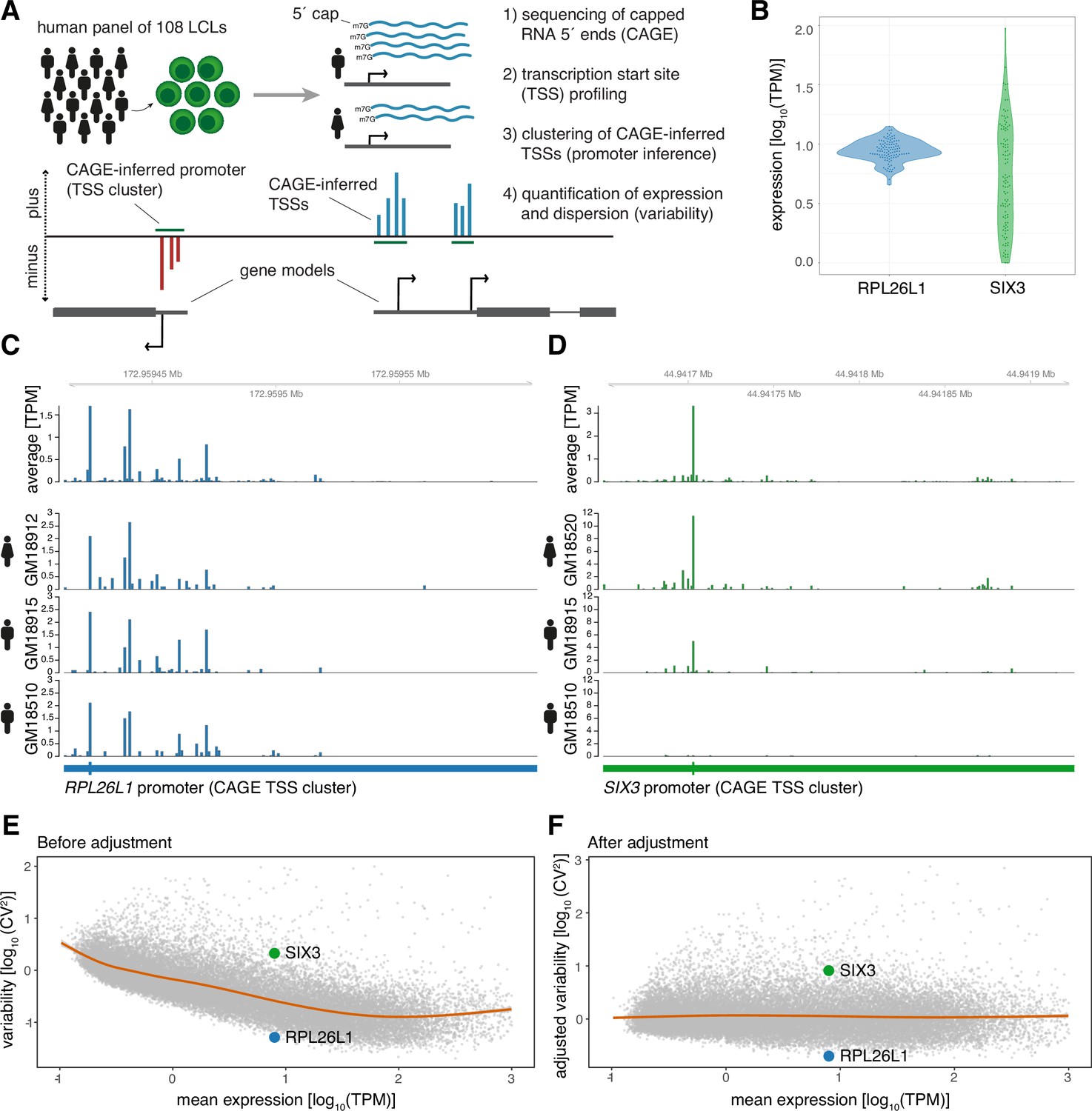
Promoter sequence and architecture determine expression variability and confer robustness to genetic variants | eLife
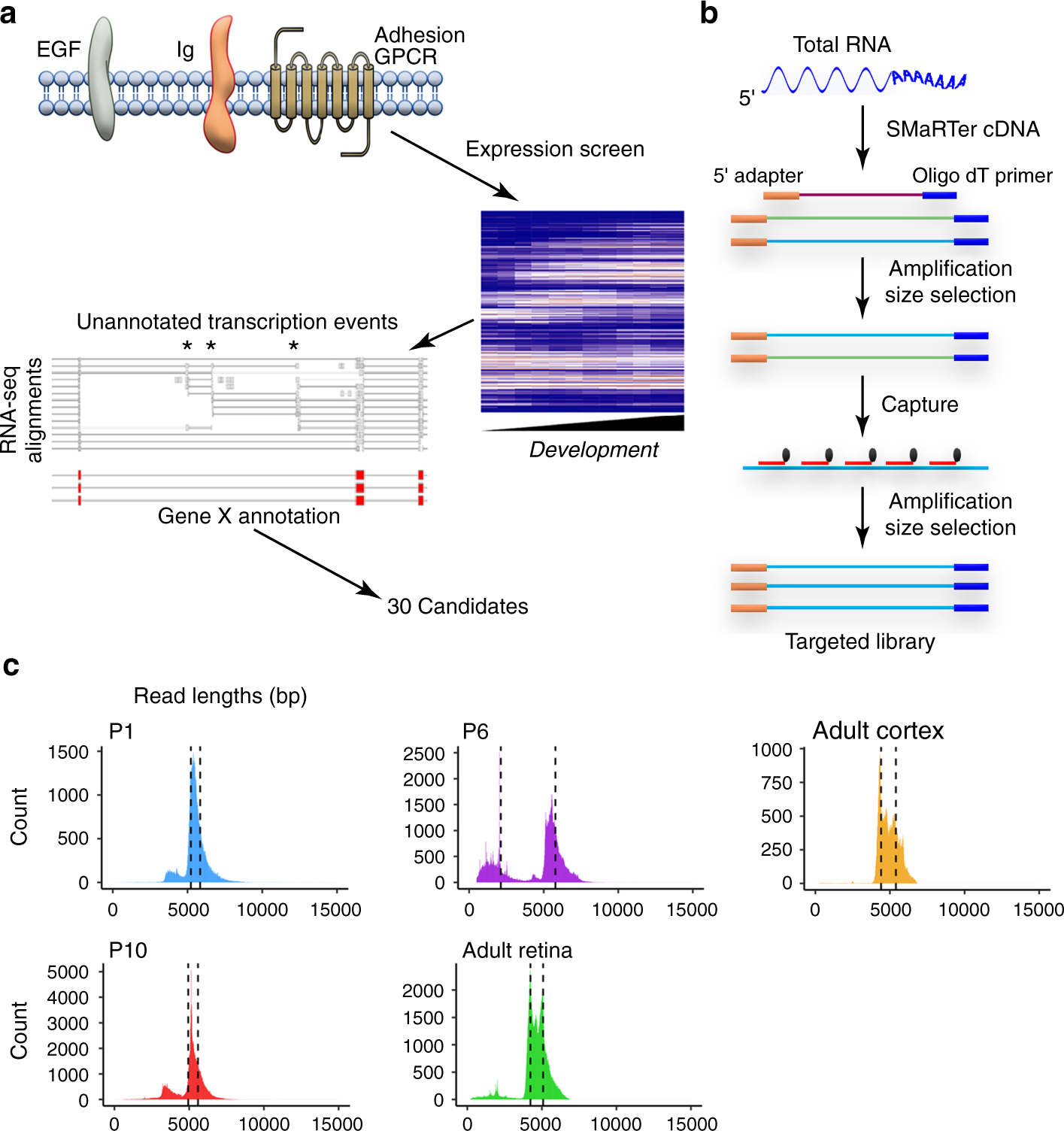
Comprehensive identification of mRNA isoforms reveals the diversity of neural cell-surface molecules with roles in retinal development and disease | Nature Communications
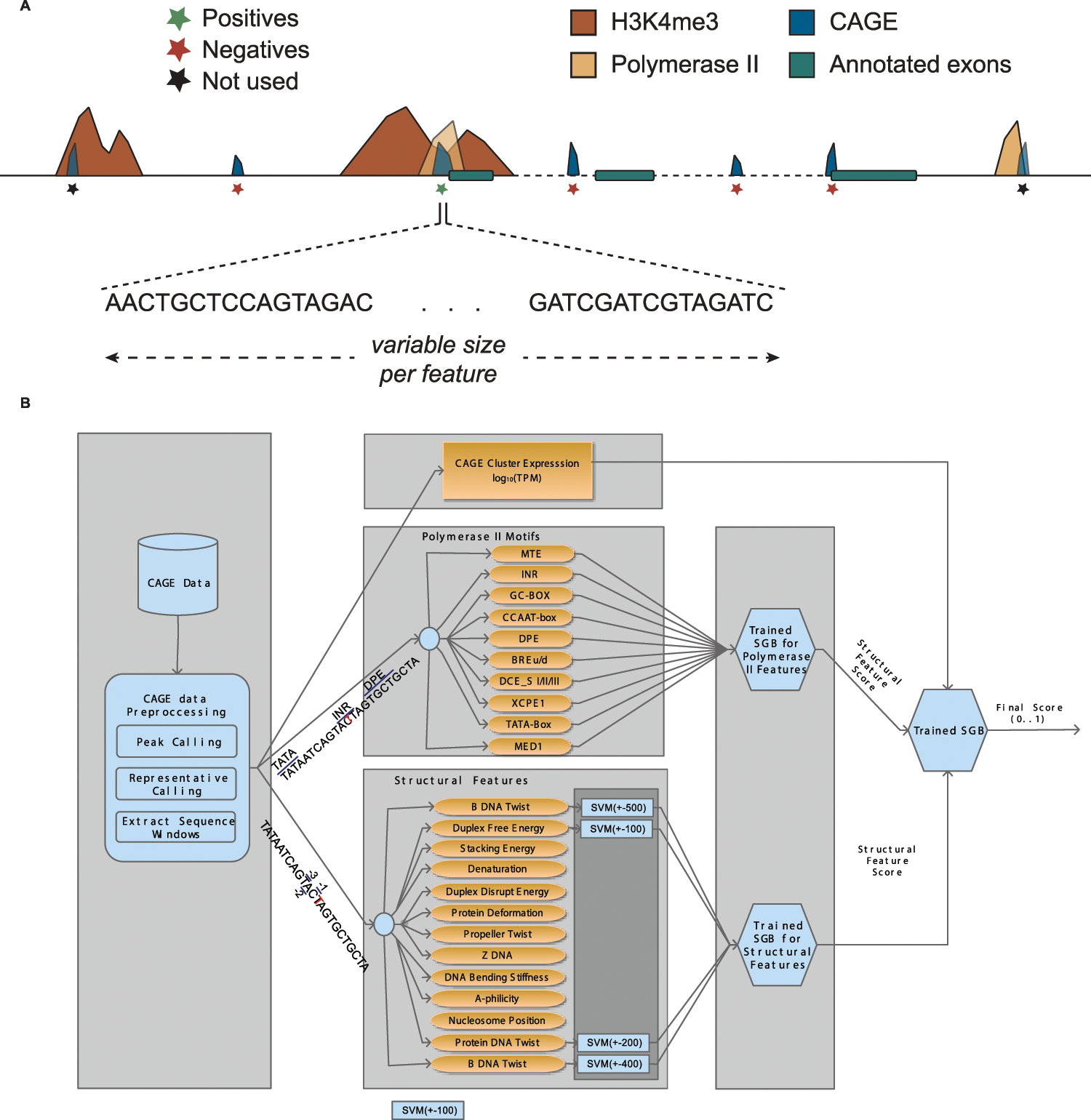
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports