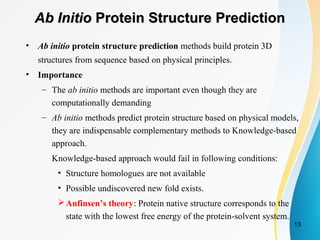
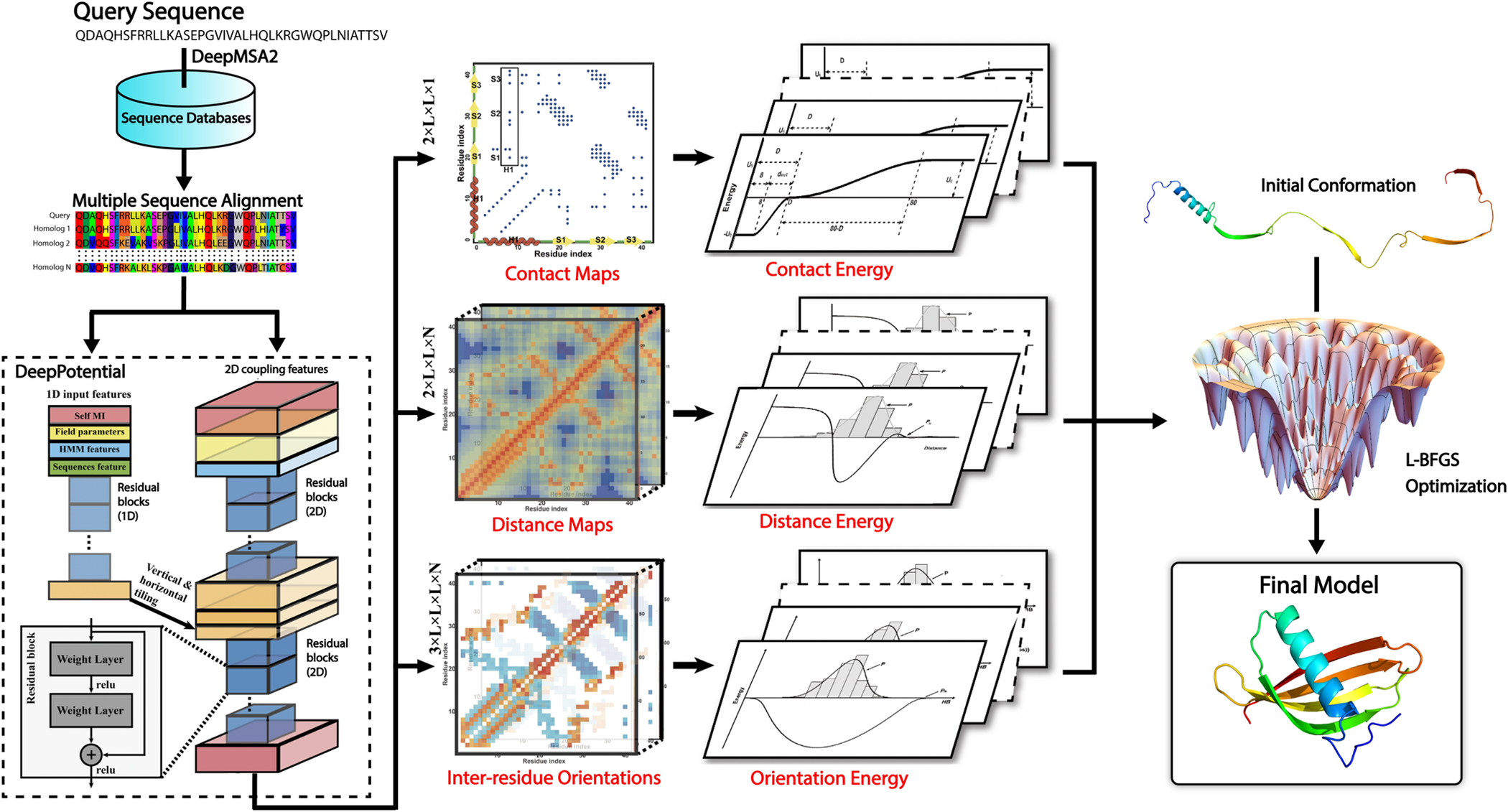
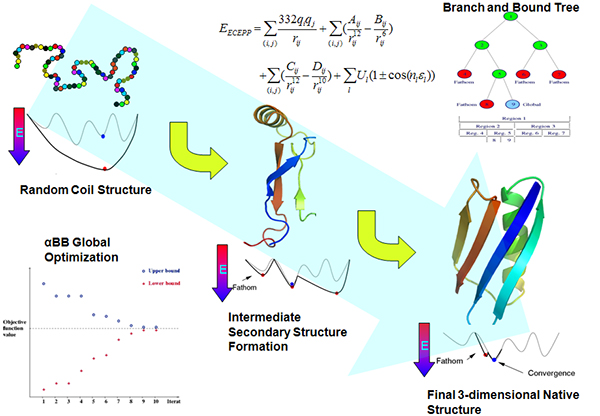
Improved fragment-based movement with LRFragLib for all-atom Ab initio protein folding | Shakhnovich Biophysics Lab

Ab initio folding of mixed-fold FSD-EY protein using formula-based polarizable hydrogen bond (PHB) charge model - ScienceDirect

Simple Model of Protein Energetics To Identify Ab Initio Folding Transitions from All-Atom MD Simulations of Proteins | Journal of Chemical Theory and Computation
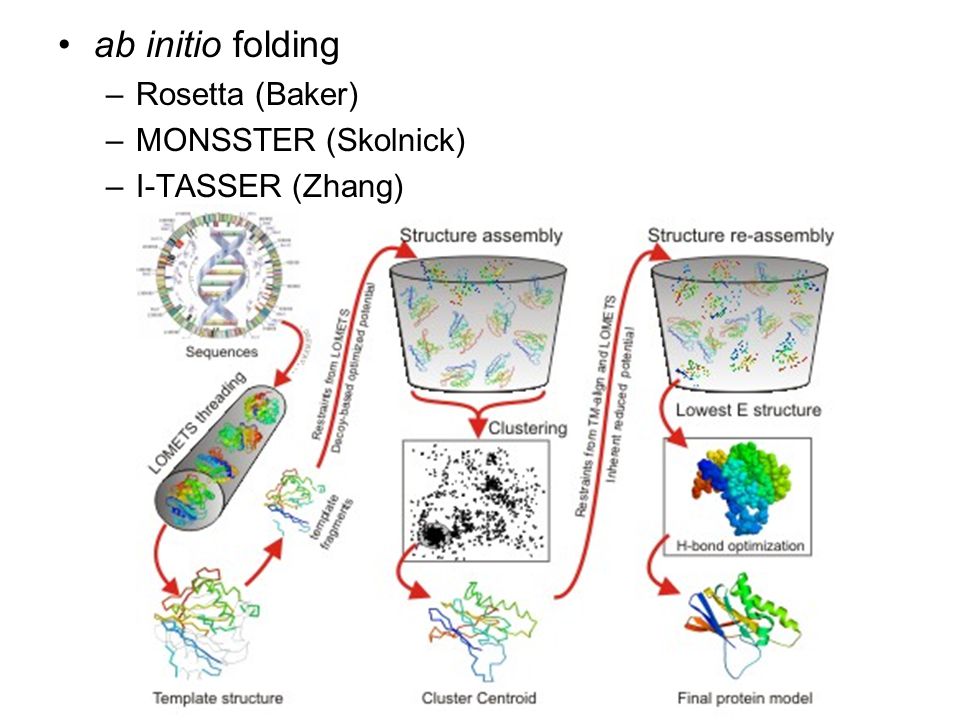
Homology Modeling comparative modeling vs. ab initio folding alignment (check gaps) threading loop building re-packing side-chains in core, DEE, SCWRL. - ppt download

Experimentally tested design coordinates and their Rosetta ab initio... | Download Scientific Diagram
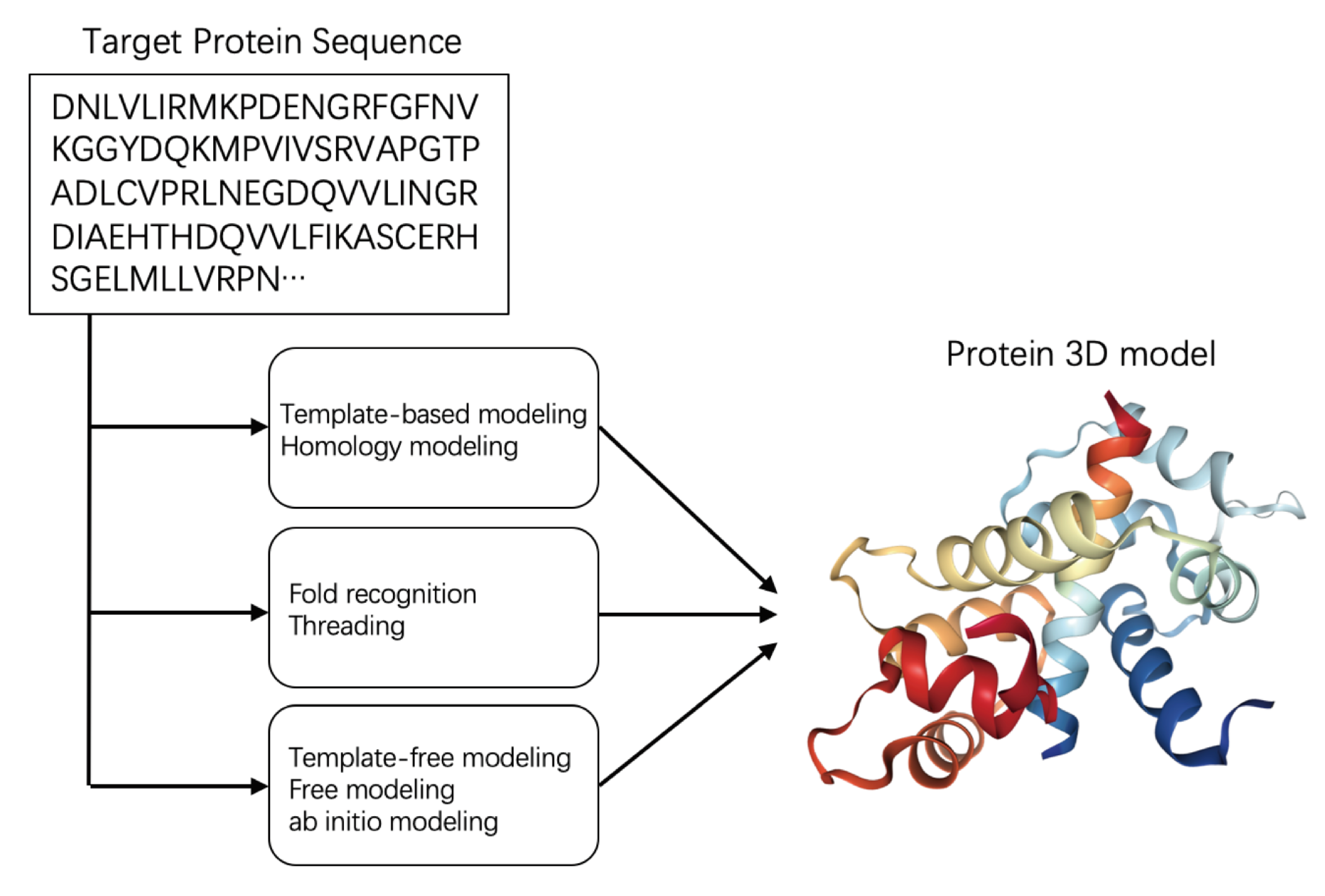
Biomolecules | Free Full-Text | Machine Learning Approaches for Quality Assessment of Protein Structures

Improved fragment sampling for ab initio protein structure prediction using deep neural networks | Nature Machine Intelligence

Threading/ Fold recognition through Phyr2 Server and Ab-initio Protein Modeling through I-TASSER - YouTube

Toward Ab Initio Protein Folding: Inherent Secondary Structure Propensity of Short Peptides from the Bioinformatics and Quantum-Chemical Perspective | The Journal of Physical Chemistry B
Automated contact distance-based ab initio protein structure prediction... | Download Scientific Diagram
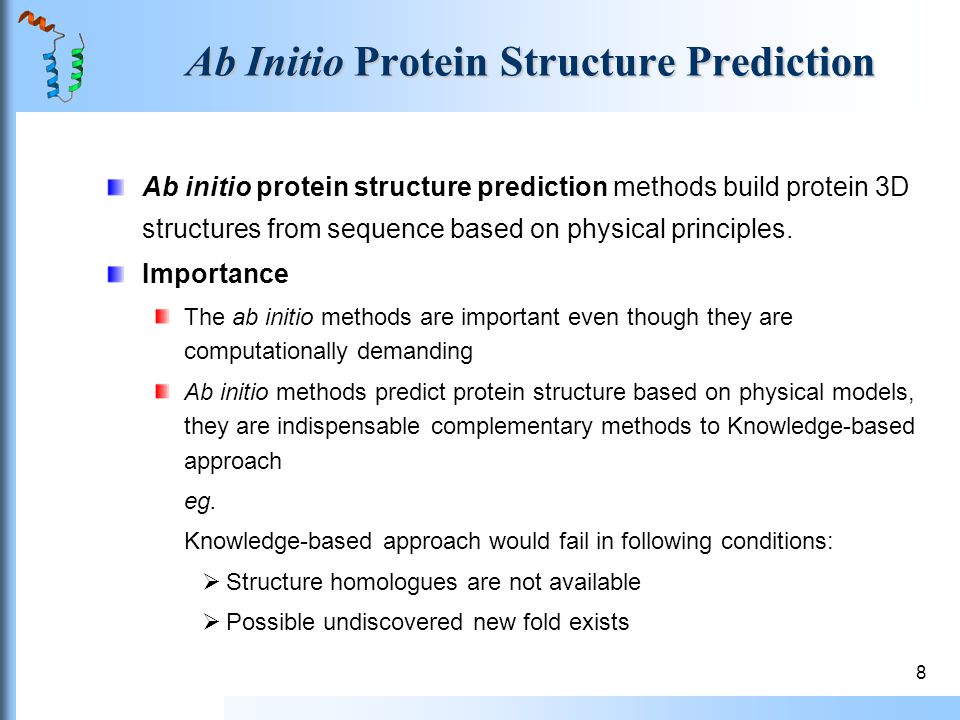
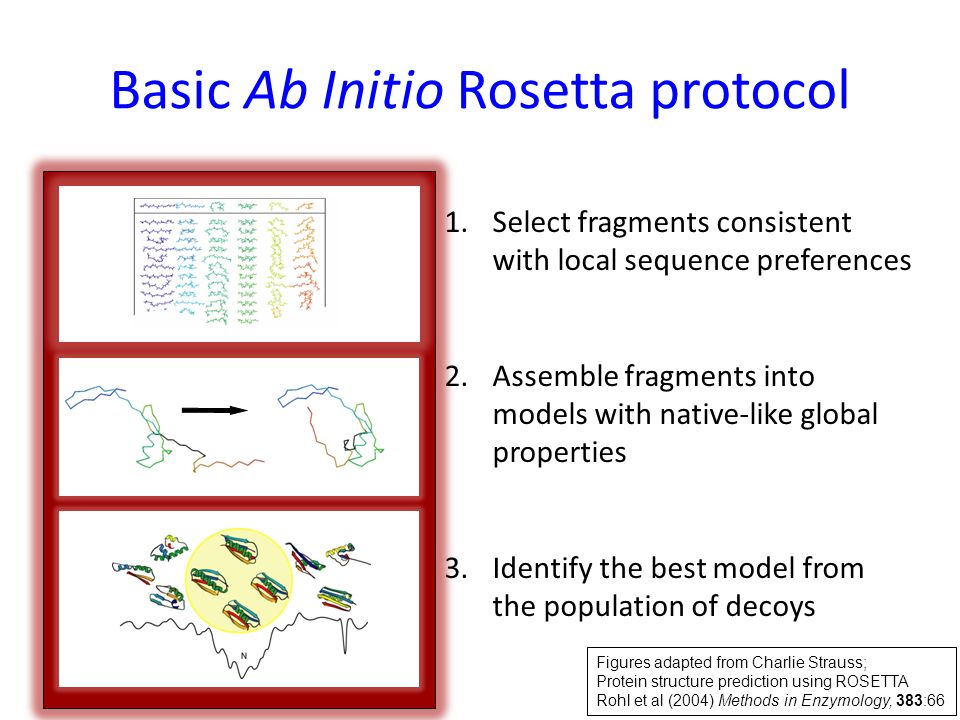





![PDF] Chapter 1 Ab Initio Protein Structure Prediction | Semantic Scholar PDF] Chapter 1 Ab Initio Protein Structure Prediction | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2d958387fa2e6183f8cd5d352cd093fc1964aa68/11-Figure1.4-1.png)


![PDF] Chapter 1 Ab Initio Protein Structure Prediction | Semantic Scholar PDF] Chapter 1 Ab Initio Protein Structure Prediction | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2d958387fa2e6183f8cd5d352cd093fc1964aa68/9-Figure1.2-1.png)